Wild genetics key to conveying potato disease resistance
New research suggests that limited genetic differences in potato lineages has left British and American spuds vulnerable to the disease that caused the Irish potato famine.
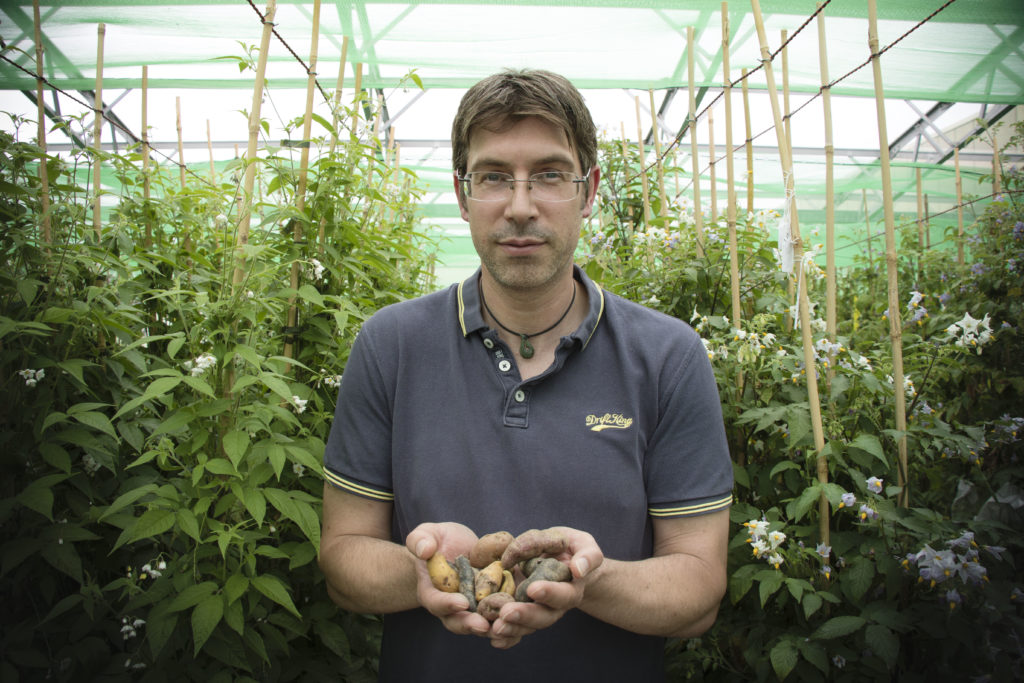
Plant scientists at the University of Dundee and the James Hutton Institute have revealed that commercial potato crops are under constant threat of late blight, the pathogen behind one of Europe’s most devastating famines. Wild genetics might provide an answer.
Ingo Hein, principal investigator in plant pathogen co-evolution, says by using tools his team has developed wild potato varieties could help breeders produce stronger, more disease-resistant varieties.
“By using the dRenSeq and PenSeq tools we have developed here in Scotland in collaboration with peers at the Sainsbury Laboratory in Norwich, we are able for the first time ever to track the historical and geographical patterns of resilient genes in American and British potatoes,” said Hein.
Their preliminary data suggests that the most commercially valuable potato varieties grown in the UK and U.S. contain a maximum of four genes that have already defeated by the late blight pathogen, Phytophthora infestans.
“Crucially, we have also been able to identify new genes that remain effective against this disease, which are not currently used in commercial potato production, and so by combining these effective genes we can prolong the longevity of individual resistances to the disease and reduce the need for chemical sprays on plants,” Hein said.
“This is highly relevant for breeding as currently the control methods for late blight in most parts of the world are based mainly on the use of chemical sprays, which can be environmentally hazardous and expensive.”
Hein was awarded £625,000 (approximately $692,000) in funding from the Biotechnology and Biological Sciences Research Council (BBSRC) to continue identifying the more-resilient potatoes. He’s working with industry partners McCain, Greenvale and James Hutton Limited.
How it works
The technology they developed came from the Potato Genome, released in 2011. Very specifically, they looked at a family of plant receptors known as nucleotide-binding, leucine-rich repeat (NLRs) genes. The researchers are interested in them because they provide resistance against many unrelated pathogens including viruses and nematodes.
“The potato genome has given us unprecedented insight into the organization of these resistances,” said Hein.
If you could write out the entire genetic code for the potato with its 840 million base pairs, it would cover a distance from Paris to Moscow. Resistance genes make up just 6 kilometers of that distance, said Hein.
“Rather than looking at 2,600 kilometers Paris to Moscow, we’re only looking at the city center of Paris, if you want an analogy,” said Hein. “So, the technology really allows us to look at these genes in particular.”
Using Resistant Gene Enrichment Sequencing (RenSeq), researchers have been able to look at the historical deployment of resistance genes. Through collaborative efforts they’ve been able to evaluate potato material that was grown in the 1800s, including the main variety grown during the famine in Ireland and Scotland. High yields made it a popular choice on marginal land. This variety and others commonly grown at the time had absolutely no resistance, said Hein.
The researchers then followed what happened after the famine years. Obviously, breeders were aware of the issue and started breeding for resistance. In the beginning, breeding was focused on seedling selection, so nothing unique was introduced through breeding at that time. It wasn’t until the 1930s that breeders realized they could use wild species to add resistance genes into new varieties.
The problem is that, since then, breeders have used the same resistance genes over and over again, giving the pathogen the ideal evolutionary battleground to adapt to these resistances. Nearly all of these resistances in the 1930s have been overcome by pathogen isolates.
The good news now is that since there are new genetic sequencing tools available, new resistance genes have been brought to market in varieties.
“That provides us with a unique opportunity to take these resistances and combine them as soon as we can so that we have a strong barrier against pathogen evolution,” said Hein. “That’s where we want to go with our project.”
More findings
The project was to officially begin in December, but preliminary data was submitted with the grant proposal. It looks promising enough that Hein received the grant.
The team of researchers has since conducted a survey of UK, Dutch and U.S. varieties, as well as some varieties from China and South Korea. They were able to quickly determine that the same resistance genes have been used over and over again, particularly in the UK and the U.S.
“There are some exceptions,” said Hein. “There are some resistances that are used in Europe that are not widely used in the U.S., and we have a little bit of evidence that the pathogen population in the U.S. has consequently changed compared to what it is in Europe.”
The project will run for three years and involves two commercial companies, McCain and Greenvale.
Some headway has already been made.
“We’ve identified some very good sources of resistances in varieties already. We have set up crosses with our own breeding companies to combine these resistances,” Hein said. “That’s going to be key. A single resistance is not going to hold out a long time.
“If we have complementary resistances combined, you stand a much better chance of having something more durable.”
— This is the latest installment of Global Perspectives, a series that looks at potato production from around the world and how international developments could affect U.S. producers. More from Global Perspectives.